Rcpp alternative to compute_centers.
Arguments
- agent
Object of the
agent-class.- a, b
Numerics denoting the parameters of the weighting function, where
ais used for the power of the function andbfor the slope of function.ais required to be positive andbshould lie between 0 and 1, where1 - bdenotes the maximal decrease in velocities in percentage.- velocities
Numeric matrix containing the change in speed for an agent whenever they move to the respective cell of this matrix.
- orientations
Numeric matrix containing the change in direction for an agent whenever they move to the respective cell of this matrix.
- time_step
Numeric denoting the number of seconds each discrete step in time should mimic. Defaults to
0.5, or half a second.
Details
Compute cell centers based on a person's current position and velocity,
accounting for potential changes in speed and direction. Alternative to
c_vd that accounts for biomechanical limitations in the
speed one can maintain when turning at a greater angle. Defaults are based on
Seethapathi et al. (2024), Brown et al. (2020), and Glaister et al. (2007).
Examples
# Create two agents, one fast and one slow
slow_agent <- agent(center = c(-2.75, 0),
radius = 0.25,
speed = 0.5,
orientation = 0,
current_goal = goal(position = c(-2.01, 0)))
fast_agent <- agent(center = c(-2.75, 0),
radius = 0.25,
speed = 2,
orientation = 0,
current_goal = goal(position = c(-2.01, 0)))
# Generate the cell centers with predped
slow_centers <- compute_centers(slow_agent,
cpp = TRUE)
fast_centers <- compute_centers(fast_agent,
cpp = TRUE)
# Generate the cell centers with m4ma
slow_m4ma <- m4ma::c_vd(1:33,
position(slow_agent),
speed(slow_agent),
orientation(slow_agent))
fast_m4ma <- m4ma::c_vd(1:33,
position(fast_agent),
speed(fast_agent),
orientation(fast_agent))
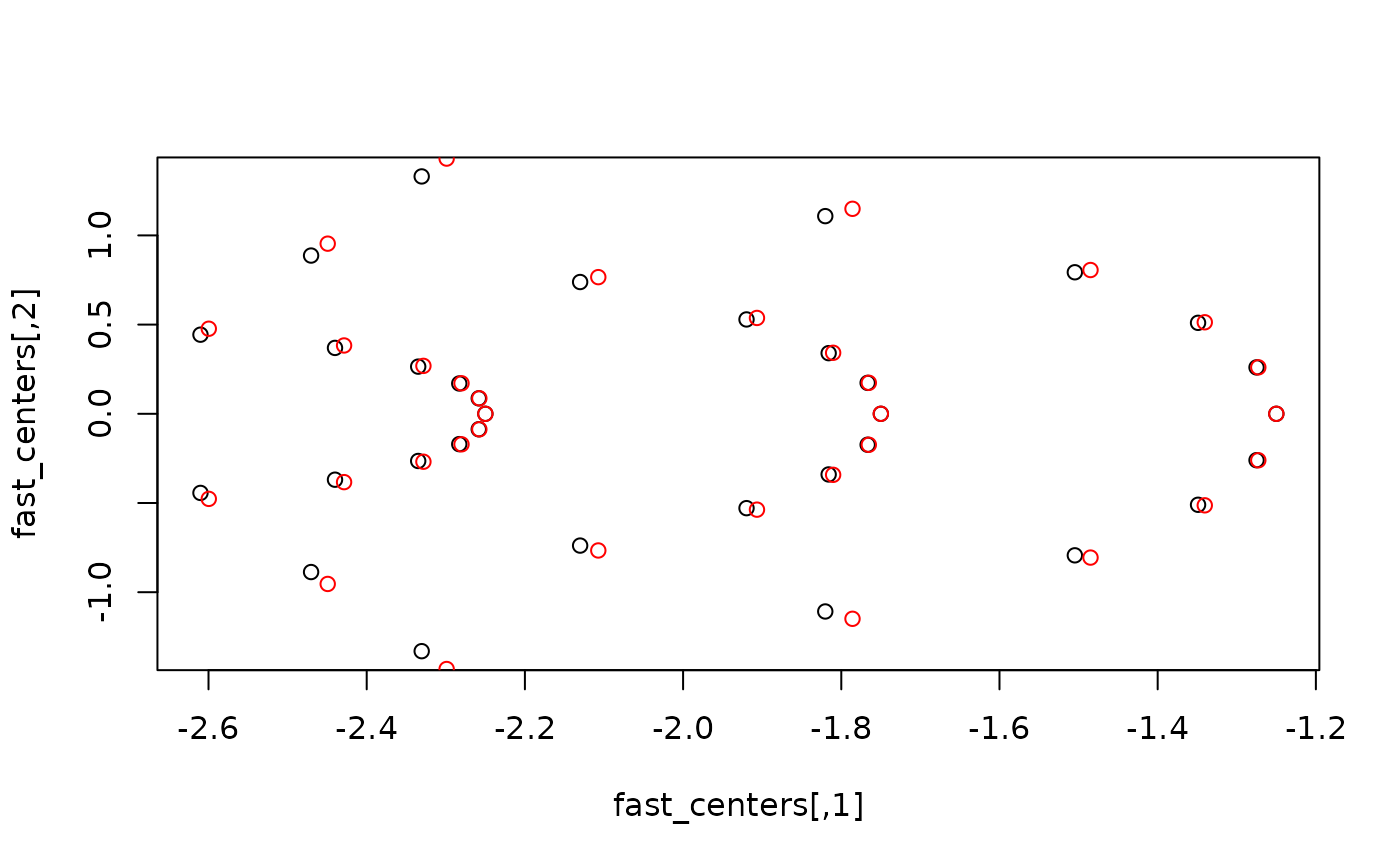
# Compare both through a plot. This should show that the predped variant
# accounts for an interaction between an agent's speed and change in
# direction when computing the cell centers
base::plot(slow_centers, col = "black")
graphics::points(slow_m4ma[, 1], slow_m4ma[, 2], col = "red")
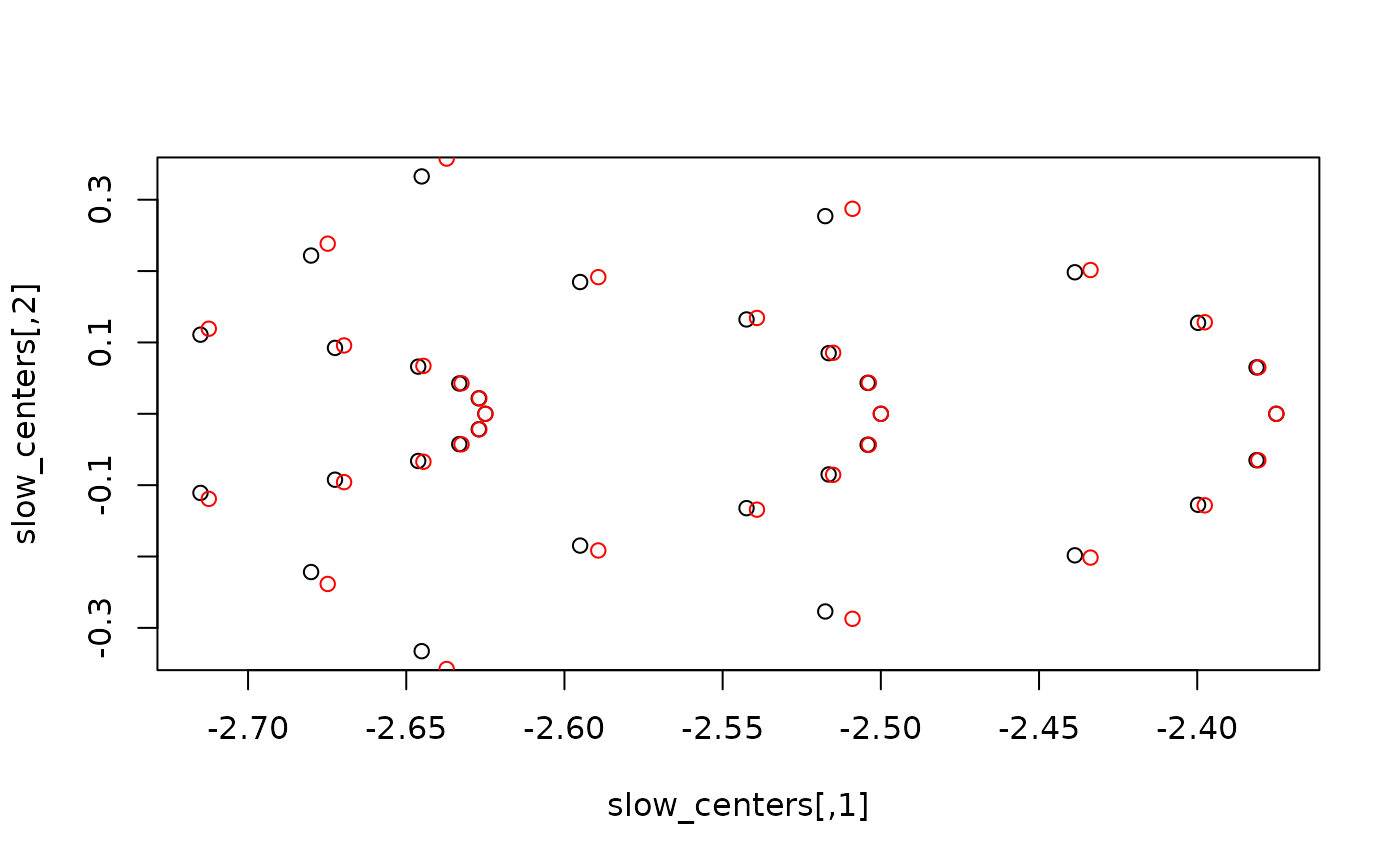 base::plot(fast_centers, col = "black")
graphics::points(fast_m4ma[, 1], fast_m4ma[, 2], col = "red")
base::plot(fast_centers, col = "black")
graphics::points(fast_m4ma[, 1], fast_m4ma[, 2], col = "red")